Submitting Raw Sequencing Read Data¶
While it is possible to submit raw sequencing read data both interactively and programmatically using similar interfaces to those used in previous exercises, it is not the recommended route. This is because files need to be uploaded to your personal dropbox area on our servers using a range of options prior to metadata submission.
We recommend using our command line interface Webin-CLI for submission of data files, as it will perform file upload automatically at time of metadata submission, making it a one step process instead of two.
For this part of the workshop, we will submit the example paired fastq files in the runs/ directory to our samples.
cd $WORKSHOP/runs/
ls *.fastq.gz
Webin-CLI¶
Webin-CLI is a Java-based utility for simultaneously uploading and submitting different types of data files. If you haven’t already done so, please see here for information on download and setup of this tool.
The main input to the Webin-CLI application is a manifest file. Included in the runs/webin-cli/ directory are manifest files for each sample. Please open paired_fastq_manifest_sample1.txt to see the format of these types of files, and what metadata is listed there. In the case of reads, this largely covers information about library preparation and sequencing platform. For information on permitted values, see here.
Note
In order to link the data files to the objects we’ve already created (study, samples), we must add the accessions received in previous steps to the manifest files.
Please replace the STUDY and SAMPLE fields with your own accessions in each file prior to submission.
First, using the -context reads setting in Webin-CLI, we will validate our submission using the -validate flag. This checks for errors in the manifest fields.
cd webin-cli/
java -jar webin-cli-4.2.0.jar -context reads -manifest paired_fastq_manifest_sample1.txt -inputDir ../ -userName user -password pass -test -validate
If this passes validation, we can replace the -validate flag with -submit to perform the submission.
Warning
Please use the -test flag to submit to our test service
java -jar webin-cli-4.2.0.jar -context reads -manifest paired_fastq_manifest_sample1.txt -inputDir ../ -userName user -password pass -test -submit
Do this for all 3 manifest files. As always, please note down the resulting run accessions (ERR numbers) for use later.
Tip
Reads have now been submitted to your samples. Feel free to move on to submitting sequences if you do not intend to use the other submission methods listed below.
Data Upload¶
Before programmatic or interactive submission, you must upload your data files (.fastq.gz in this case) to your dropbox area on our servers. In order to check for successful file transfers, we must compute MD5s for each file prior to upload. For example:
for s in SARS-CoV-2-Sample*
> do
> md5 $s
> done
MD5 (SARS-CoV-2-Sample1_1.fastq.gz) = a5e42219e299c1a0bcadd2b67bf7b32d
MD5 (SARS-CoV-2-Sample1_2.fastq.gz) = df4480daa2e9b4c2dfa7c5384b281e11
MD5 (SARS-CoV-2-Sample2_1.fastq.gz) = 75c798a4f452ef5cb1ee7ab0c2df250c
MD5 (SARS-CoV-2-Sample2_2.fastq.gz) = cbc484055e1c82814d62e8dc62088f19
MD5 (SARS-CoV-2-Sample3_1.fastq.gz) = 29e51011d86ad0c8fc8681d50136dffc
MD5 (SARS-CoV-2-Sample3_2.fastq.gz) = 9cb639082ebf3c3172ba595602b3f07e
Next, select the most convenient upload option for your system from our range of options. Upload all 3 pairs of files.
Programmatic¶
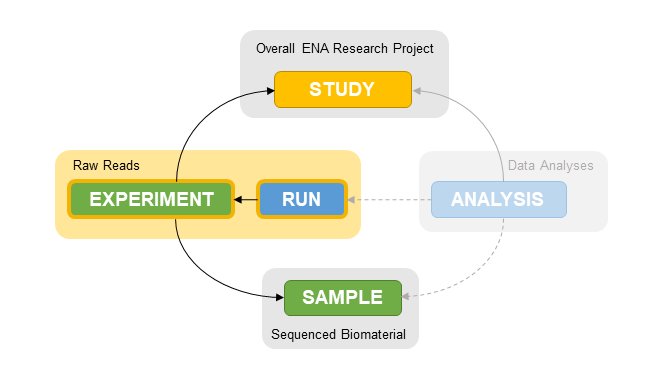
As in previous steps, this type of submission is performed using XML files. In the case of reads, we must submit 2 types of object: experiments and reads. Experiments hold information about library preparation and sequencing protocols, and link to studies and samples. Runs simply link experiments and data files.
Please find example XMLs for both experiments and reads in the runs/programmatic directory and make the following edits:
in
experiments.xml, replace all occurrences ofPRJEB####with your study accession, and all occurrences ofSAME######with the equivalent sample accessions.in
runs.xml, replace thechecksumfield in each<FILE>tag with those that you computed earlier. These will be used to check for file corruption that may have occurred during upload.
Note that runs reference experiments by their aliases. E.g.
in
experiments.xml:<EXPERIMENT alias="SARS-CoV-2 experiment 1">in
runs.xml:<EXPERIMENT_REF refname="SARS-CoV-2 experiment 1" />
Note
In all object referencing/linking (using <*_REF> tags), we can link by either alias, using refname="" or by accession, using accession=""
As with previous programmatic submissions, we will send these XML files to our test service using cURL:
curl -u username:password -F "SUBMISSION=@submission.xml" -F "EXPERIMENT=@experiments.xml" -F "RUN=@runs.xml" "https://wwwdev.ebi.ac.uk/ena/submit/drop-box/submit/"
Interactive¶
Runs can be created through the Webin Submissions Portal using a tab-delimited (.tsv) spreadsheet. Please find an example spreadsheet in the runs/interactive/ folder.
In this spreadsheet, there are several fields you will need to update in order to correctly link and validate your data files.
replace all occurrences of
ERS#######with your equivalent sample accessionsreplace all occurrences of
PRJEB#####with your equivalent study accessionreplace both
forward_file_md5andreverse_file_md5fields with the hashes you calculated earlier - these will be used to check for file corruption during upload
In the Webin Submissions Portal dashboard, the reads menu is colour coded in orange. Choose the ‘Submit Reads’ button and upload the spreadsheet.
Tip
Reads have now been submitted to your samples. Let’s move on to submitting sequences.